Our lab develops and uses cutting-edge techniques to study the physics of proteins under force. We combine single molecule force spectroscopy and bio-materials design with protein engineering, surface chemistry and computer programming. Our long-term goal is to understand how proteins operating under force use their folding state as a mechanical signal, and how this signal propagates from single molecule level to tissue and organs.
Movie shows a titin domain in series with a spring unfolding and refolding under force; part of SI of our J. Chem. Educ. 2022 article.
1. SINGLE MOLECULE NANO-MECHANICS
All processes in the human body rely to a large extent on the correct functioning of proteins. To perform their intended function, proteins need to fold in a specific 3D structure. Protein folding is one of the most complex physical process that combines short ranged hydrogen bonds between amino acids with the entropy of the polypeptide chain and interactions with the surrounding solvent molecules.
In our laboratory we use a revolutionary single molecule technique, based on magnetic tweezers and covalent attachment. This technique can expose single protein constructs to mechanical perturbations identical to those experienced in vivo, and for extended time, of up to 15 days per molecule.Using single molecule magnetic tweezers, we can directly sample the folding and unfolding process of proteins using force as a perturbation. We measure the end-to-end length of multi-domain protein constructs, which show under force unfolding and refolding transitions as step increments.
We are currently working together with Vali Raicu from UWM on developing a FRET/magnetic tweezers setup that would add a new dimension to our measurements. Using fluorescent tags, we plan to monitor the binding and unbinding of different reaction partners to single proteins under force.
2. BIO-INSPIRED ENGINEERING
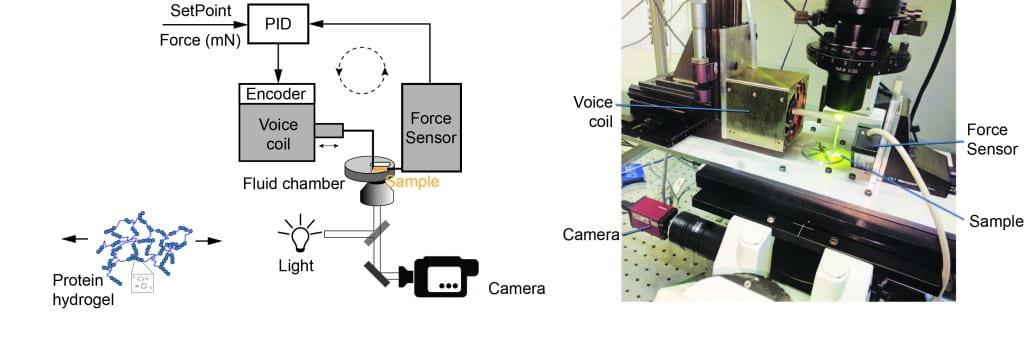
Another direction of our laboratory is to develop “smart” bio-inspired materials, with possible biomedical applications for tissue engineering, drug delivery, bio-sensors and bio-robotics. We use chemical cross-linking of engineered multi-protein constructs to synthesize protein hydrogels. These gels have remarkable properties and follow the measured unfolding mechanics of proteins measured with single molecule techniques. We have constructed two instruments which allow us to measure the mechanical response of protein hydrogels under force-clamp conditions. Combining this technique with optical microscopy will allow us to monitor the complete structural transformation of polyprotein hydrogels. To trigger biological-like motion, we combine these protein hydrogels with small molecules or polymers, which allow us to program them in new temporary shapes. To induce shape changes, we then induce a transition in either the protein primary network or the small molecules/polymeric secondary network.